I have training (X) and test data (test_data_process) set with the same columns and order, as indicated below:
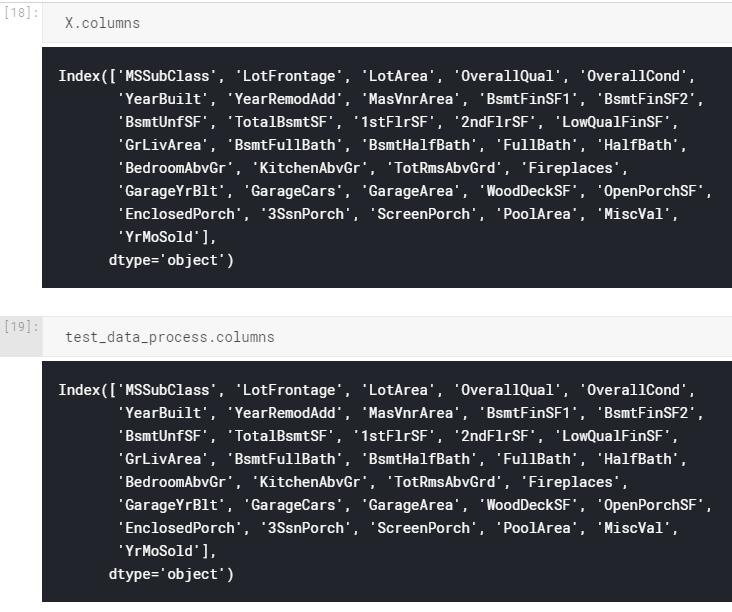
But when I do
predictions = my_model.predict(test_data_process)
It gives the following error:
ValueError: feature_names mismatch: ['f0', 'f1', 'f2', 'f3', 'f4', 'f5', 'f6', 'f7', 'f8', 'f9', 'f10', 'f11', 'f12', 'f13', 'f14', 'f15', 'f16', 'f17', 'f18', 'f19', 'f20', 'f21', 'f22', 'f23', 'f24', 'f25', 'f26', 'f27', 'f28', 'f29', 'f30', 'f31', 'f32', 'f33', 'f34'] ['MSSubClass', 'LotFrontage', 'LotArea', 'OverallQual', 'OverallCond', 'YearBuilt', 'YearRemodAdd', 'MasVnrArea', 'BsmtFinSF1', 'BsmtFinSF2', 'BsmtUnfSF', 'TotalBsmtSF', '1stFlrSF', '2ndFlrSF', 'LowQualFinSF', 'GrLivArea', 'BsmtFullBath', 'BsmtHalfBath', 'FullBath', 'HalfBath', 'BedroomAbvGr', 'KitchenAbvGr', 'TotRmsAbvGrd', 'Fireplaces', 'GarageYrBlt', 'GarageCars', 'GarageArea', 'WoodDeckSF', 'OpenPorchSF', 'EnclosedPorch', '3SsnPorch', 'ScreenPorch', 'PoolArea', 'MiscVal', 'YrMoSold'] expected f22, f25, f0, f34, f32, f5, f20, f3, f33, f15, f24, f31, f28, f9, f8, f19, f14, f18, f17, f2, f13, f4, f27, f16, f1, f29, f11, f26, f10, f7, f21, f30, f23, f6, f12 in input data training data did not have the following fields: OpenPorchSF, BsmtFinSF1, LotFrontage, GrLivArea, YrMoSold, FullBath, TotRmsAbvGrd, GarageCars, YearRemodAdd, BedroomAbvGr, PoolArea, KitchenAbvGr, LotArea, HalfBath, MiscVal, EnclosedPorch, BsmtUnfSF, MSSubClass, BsmtFullBath, YearBuilt, 1stFlrSF, ScreenPorch, 3SsnPorch, TotalBsmtSF, GarageYrBlt, MasVnrArea, OverallQual, Fireplaces, WoodDeckSF, 2ndFlrSF, BsmtFinSF2, BsmtHalfBath, LowQualFinSF, OverallCond, GarageArea
So it complains that the training data (X) does not have those fields, whereas it has.
How to solve this issue?
[UPDATE]:
My code:
X = data.select_dtypes(exclude=['object']).drop(columns=['Id'])
X['YrMoSold'] = X['YrSold'] * 12 + X['MoSold']
X = X.drop(columns=['YrSold', 'MoSold', 'SalePrice'])
X = X.fillna(0.0000001)
train_X, val_X, train_y, val_y = train_test_split(X.values, y.values, test_size=0.2)
my_model = XGBRegressor(n_estimators=100, learning_rate=0.05, booster='gbtree')
my_model.fit(train_X, train_y, early_stopping_rounds=5,
eval_set=[(val_X, val_y)], verbose=False)
test_data_process = test_data.select_dtypes(exclude=['object']).drop(columns=['Id'])
test_data_process['YrMoSold'] = test_data_process['YrSold'] * 12 + test_data['MoSold']
test_data_process = test_data_process.drop(columns=['YrSold', 'MoSold'])
test_data_process = test_data_process.fillna(0.0000001)
test_data_process = test_data_process[X.columns]
predictions = my_model.predict(test_data_process)






 已为社区贡献126445条内容
已为社区贡献126445条内容

所有评论(0)